Challenge #4
Click image to enlarge
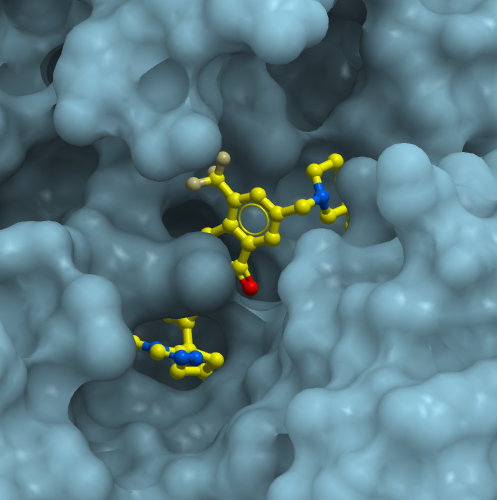
Finding ligands targeting the TKB domain of CBLB
The fourth CACHE challenge is focused on the TKB domain of CBLB, an E3 ubiquitin-protein ligase with hundreds of compounds reported in the patent literature from multiple organizations, all chemically related. We provide an SD file with 895 of these compounds and IC50’s ranging from low nM to low µM. A crystal structure (PDB code 8GCY) with one of these compounds reveals that the inhibitor exploits a closed/inactive conformational state of the protein. CACHE participants are asked to predict compounds that (1) bind the same pocket and compete with the co-crystallized compound, (2) represent novel chemical templates and (3) bind with a KD below 30 micromolar. Read more under Details below.
CACHE will cover 50% of the costs of testing compounds experimentally, estimated at $1,000, (including attempts to co-crystallize the best compounds by the team at the SGC that solved structure 8GCY). In addition, CACHE will cover 50% of the costs of procuring the compounds (capped at $10,000) for academics and companies with less than 20 employees, and 75% of the costs for participants who make their software opensource.
A total of 25 applicants will be invited to participate to this challenge, following a double-blind peer review process where each applicant is asked to review 5 applications.
CACHE is strongly committed to equity, diversity and inclusion. We strive to create an initiative that welcomes participants from countries, institutions and groups that are underrepresented in our research community. We endeavour to work with individuals who contribute to furthering the diversity of ideas and approaches, welcoming all communities to participate in our competitions.
All applications and submissions to this CACHE Challenge are subject to the CACHE Terms of Participation. Applications should be submitted using the online application form.
2023-03-09
2023-05-09
Timeline

Details
TKB domain of CBLB
| Parameter | Description | |
|---|---|---|
| Accession |
Q13191 · CBLB_HUMAN |
|
| Disease association | Several cancers (PMID: 33306199), potential immunotherapy (PMID: 24875217), in phase one clinical trials (NX-1607). | |
| Challenge | Predict compounds that bind to the closed conformation of the CBLB TKB domain with novel chemical templates and KD below 30 micromolar. | |
| Why the TKB domain? | Compounds binding to the TKB domain induce a closed confirmation as observed in the provided co-crystal structure (PDB code: 8GCY) and inactivate CBLB while stabilizing it (significant thermal shift by DSF, unpublished data). Compounds with similar structural mechanism have been shown to be active in several preclinical cancer models ( https://www.nurixtx.com/wp-content/uploads/2022/09/Nurix-CBL-B-DOT-Talk_JK.pdf) and NX-1607, a compound with the same core structure, is in phase 1 clinical trials for multiple oncology indications. | |
| Pharmacological landscape | Several potent small molecule inhibitors of CBLB have been reported in patent applications (WO2019148005, WO2021021761, WO2020210508, WO2020236654, WO2020264398) | |
| Macromolecular structures | The SGC has reported a 2 Å resolution co-crystal structure of a potent CBLB inhibitor from the patent literature in complex with the closed/inactive CBLB TKB domain. Structures of CBLB in its open/active state (e.g. 3ZNI, 3PFV) are not relevant to this challenge. | |
